Fluorine »
PDB 2pdl-2q94 »
2q2y »
Fluorine in PDB 2q2y: Crystal Structure of Ksp in Complex with Inhibitor 1
Protein crystallography data
The structure of Crystal Structure of Ksp in Complex with Inhibitor 1, PDB code: 2q2y
was solved by
Y.Yan,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 39.90 / 2.50 |
| Space group | P 21 21 21 |
| Cell size a, b, c (Å), α, β, γ (°) | 68.860, 79.800, 159.570, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 19 / 25.2 |
Other elements in 2q2y:
The structure of Crystal Structure of Ksp in Complex with Inhibitor 1 also contains other interesting chemical elements:
| Magnesium | (Mg) | 2 atoms |
Fluorine Binding Sites:
The binding sites of Fluorine atom in the Crystal Structure of Ksp in Complex with Inhibitor 1
(pdb code 2q2y). This binding sites where shown within
5.0 Angstroms radius around Fluorine atom.
In total 2 binding sites of Fluorine where determined in the Crystal Structure of Ksp in Complex with Inhibitor 1, PDB code: 2q2y:
Jump to Fluorine binding site number: 1; 2;
In total 2 binding sites of Fluorine where determined in the Crystal Structure of Ksp in Complex with Inhibitor 1, PDB code: 2q2y:
Jump to Fluorine binding site number: 1; 2;
Fluorine binding site 1 out of 2 in 2q2y
Go back to
Fluorine binding site 1 out
of 2 in the Crystal Structure of Ksp in Complex with Inhibitor 1
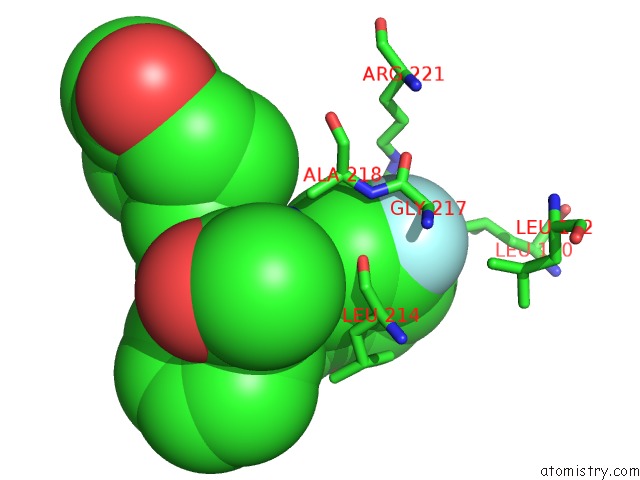
Mono view
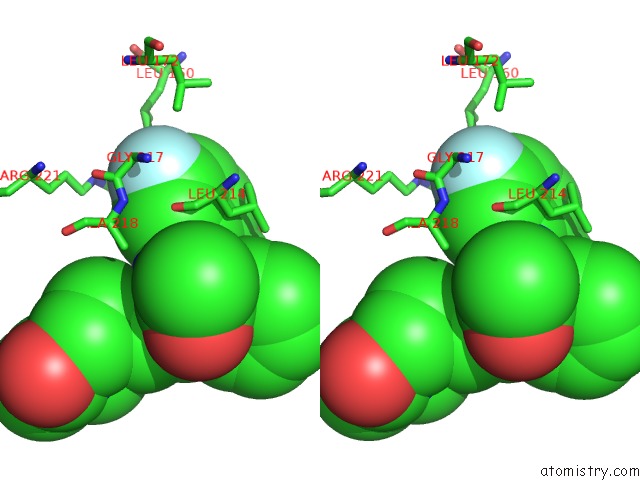
Stereo pair view

Mono view

Stereo pair view
A full contact list of Fluorine with other atoms in the F binding
site number 1 of Crystal Structure of Ksp in Complex with Inhibitor 1 within 5.0Å range:
|
Fluorine binding site 2 out of 2 in 2q2y
Go back to
Fluorine binding site 2 out
of 2 in the Crystal Structure of Ksp in Complex with Inhibitor 1
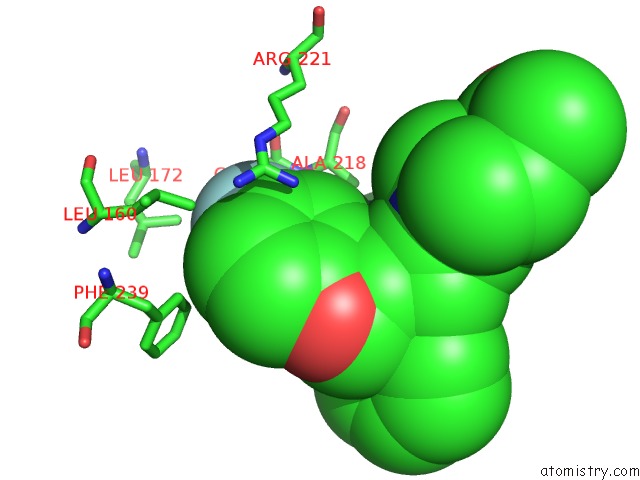
Mono view
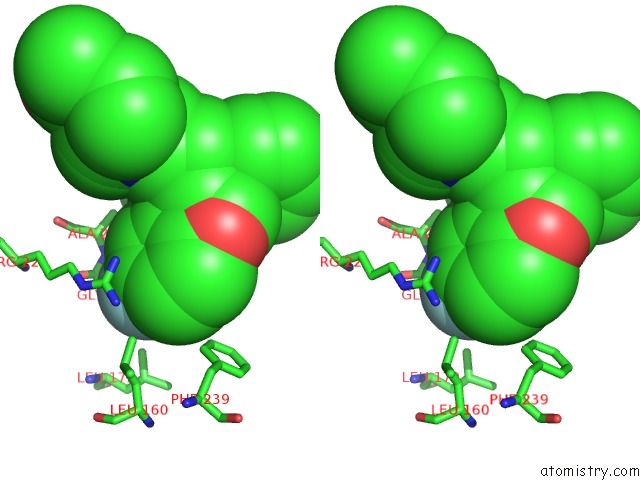
Stereo pair view

Mono view

Stereo pair view
A full contact list of Fluorine with other atoms in the F binding
site number 2 of Crystal Structure of Ksp in Complex with Inhibitor 1 within 5.0Å range:
|
Reference:
A.J.Roecker,
P.J.Coleman,
S.P.Mercer,
J.D.Schreier,
C.A.Buser,
E.S.Walsh,
K.Hamilton,
R.B.Lobell,
W.Tao,
R.E.Diehl,
V.J.South,
J.P.Davide,
N.E.Kohl,
Y.Yan,
L.C.Kuo,
C.Li,
C.Fernandez-Metzler,
E.A.Mahan,
T.Prueksaritanont,
G.D.Hartman.
Kinesin Spindle Protein (Ksp) Inhibitors. Part 8: Design and Synthesis of 1,4-Diaryl-4,5-Dihydropyrazoles As Potent Inhibitors of the Mitotic Kinesin Ksp. Bioorg.Med.Chem.Lett. V. 17 5677 2007.
ISSN: ISSN 0960-894X
PubMed: 17766111
DOI: 10.1016/J.BMCL.2007.07.074
Page generated: Mon Jul 14 14:04:51 2025
ISSN: ISSN 0960-894X
PubMed: 17766111
DOI: 10.1016/J.BMCL.2007.07.074
Last articles
F in 7HGPF in 7HH1
F in 7H6M
F in 7HG5
F in 7HG7
F in 7H9A
F in 7HBL
F in 7HB6
F in 7HAP
F in 7H7I