Fluorine »
PDB 3fls-3g70 »
3fzr »
Fluorine in PDB 3fzr: Crystal Structure of PYK2 Complexed with Pf-431396
Enzymatic activity of Crystal Structure of PYK2 Complexed with Pf-431396
All present enzymatic activity of Crystal Structure of PYK2 Complexed with Pf-431396:
2.7.10.2;
2.7.10.2;
Protein crystallography data
The structure of Crystal Structure of PYK2 Complexed with Pf-431396, PDB code: 3fzr
was solved by
S.Han,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 17.88 / 2.70 |
| Space group | P 4 21 2 |
| Cell size a, b, c (Å), α, β, γ (°) | 107.326, 107.326, 75.786, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 21.1 / 27.4 |
Fluorine Binding Sites:
The binding sites of Fluorine atom in the Crystal Structure of PYK2 Complexed with Pf-431396
(pdb code 3fzr). This binding sites where shown within
5.0 Angstroms radius around Fluorine atom.
In total 3 binding sites of Fluorine where determined in the Crystal Structure of PYK2 Complexed with Pf-431396, PDB code: 3fzr:
Jump to Fluorine binding site number: 1; 2; 3;
In total 3 binding sites of Fluorine where determined in the Crystal Structure of PYK2 Complexed with Pf-431396, PDB code: 3fzr:
Jump to Fluorine binding site number: 1; 2; 3;
Fluorine binding site 1 out of 3 in 3fzr
Go back to
Fluorine binding site 1 out
of 3 in the Crystal Structure of PYK2 Complexed with Pf-431396
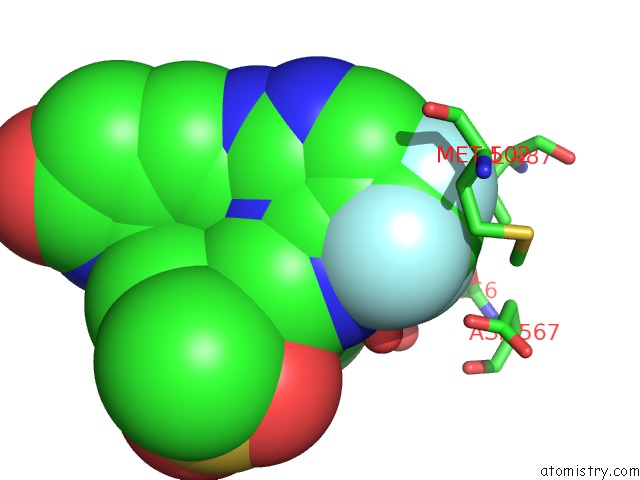
Mono view
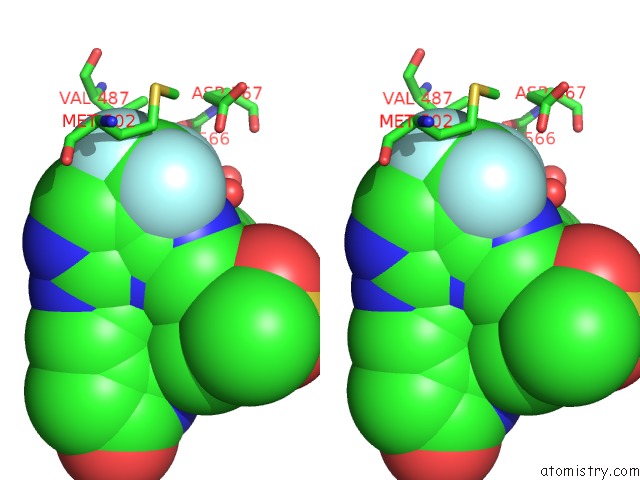
Stereo pair view

Mono view

Stereo pair view
A full contact list of Fluorine with other atoms in the F binding
site number 1 of Crystal Structure of PYK2 Complexed with Pf-431396 within 5.0Å range:
|
Fluorine binding site 2 out of 3 in 3fzr
Go back to
Fluorine binding site 2 out
of 3 in the Crystal Structure of PYK2 Complexed with Pf-431396
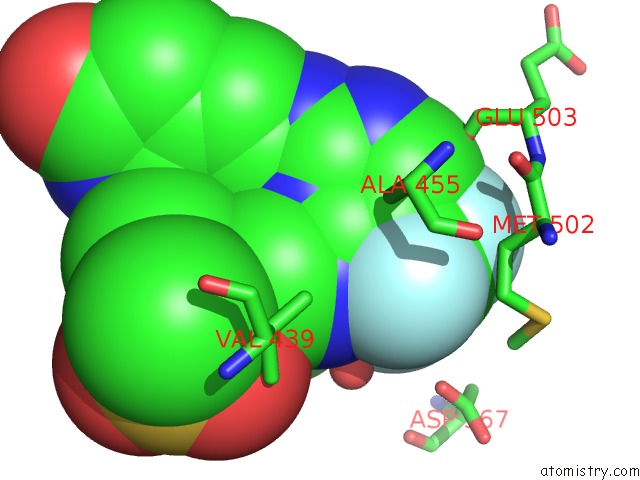
Mono view
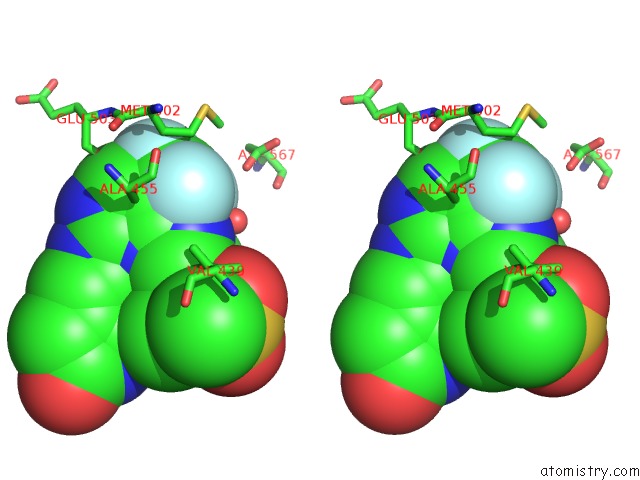
Stereo pair view

Mono view

Stereo pair view
A full contact list of Fluorine with other atoms in the F binding
site number 2 of Crystal Structure of PYK2 Complexed with Pf-431396 within 5.0Å range:
|
Fluorine binding site 3 out of 3 in 3fzr
Go back to
Fluorine binding site 3 out
of 3 in the Crystal Structure of PYK2 Complexed with Pf-431396
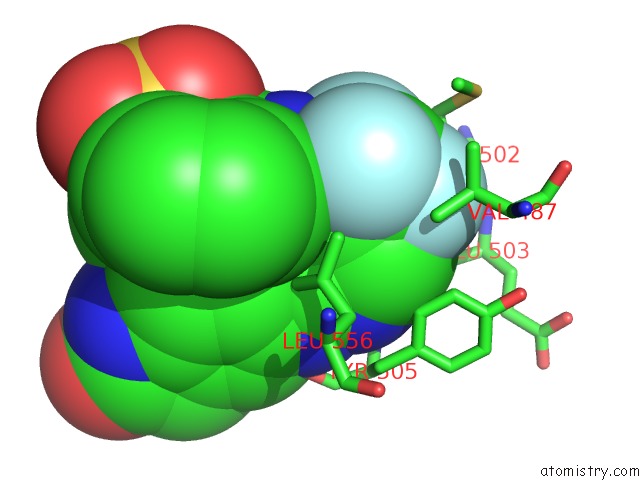
Mono view
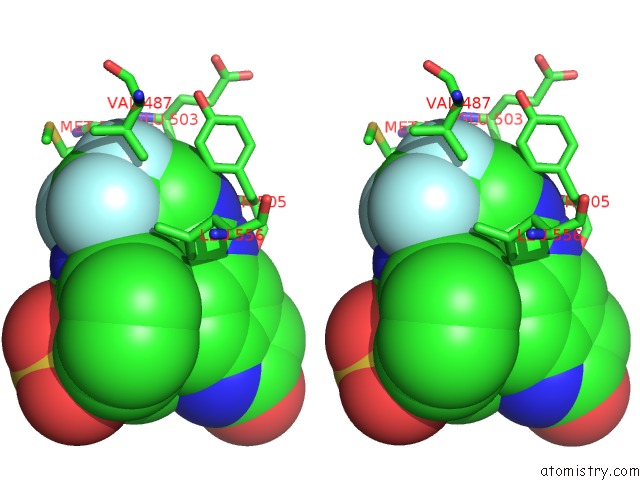
Stereo pair view

Mono view

Stereo pair view
A full contact list of Fluorine with other atoms in the F binding
site number 3 of Crystal Structure of PYK2 Complexed with Pf-431396 within 5.0Å range:
|
Reference:
S.Han,
A.Mistry,
J.S.Chang,
D.Cunningham,
M.Griffor,
P.C.Bonnette,
H.Wang,
B.A.Chrunyk,
G.E.Aspnes,
D.P.Walker,
A.D.Brosius,
L.Buckbinder.
Structural Characterization of Proline-Rich Tyrosine Kinase 2 (PYK2) Reveals A Unique (Dfg-Out) Conformation and Enables Inhibitor Design. J.Biol.Chem. V. 284 13193 2009.
ISSN: ISSN 0021-9258
PubMed: 19244237
DOI: 10.1074/JBC.M809038200
Page generated: Mon Jul 14 16:24:28 2025
ISSN: ISSN 0021-9258
PubMed: 19244237
DOI: 10.1074/JBC.M809038200
Last articles
F in 7JTOF in 7JOM
F in 7JT4
F in 7JRA
F in 7JR6
F in 7JPW
F in 7JPL
F in 7JPK
F in 7JL3
F in 7JN3