Fluorine »
PDB 4zga-4zzi »
4zx4 »
Fluorine in PDB 4zx4: X-Ray Crystal Structure of Pfa-M1 in Complex with Hydroxamic Acid- Based Inhibitor 10O
Protein crystallography data
The structure of X-Ray Crystal Structure of Pfa-M1 in Complex with Hydroxamic Acid- Based Inhibitor 10O, PDB code: 4zx4
was solved by
N.Drinkwater,
S.Mcgowan,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 34.04 / 1.90 |
| Space group | P 21 21 21 |
| Cell size a, b, c (Å), α, β, γ (°) | 75.150, 109.190, 118.830, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 15.8 / 19.7 |
Other elements in 4zx4:
The structure of X-Ray Crystal Structure of Pfa-M1 in Complex with Hydroxamic Acid- Based Inhibitor 10O also contains other interesting chemical elements:
| Magnesium | (Mg) | 1 atom |
| Zinc | (Zn) | 1 atom |
Fluorine Binding Sites:
The binding sites of Fluorine atom in the X-Ray Crystal Structure of Pfa-M1 in Complex with Hydroxamic Acid- Based Inhibitor 10O
(pdb code 4zx4). This binding sites where shown within
5.0 Angstroms radius around Fluorine atom.
In total 3 binding sites of Fluorine where determined in the X-Ray Crystal Structure of Pfa-M1 in Complex with Hydroxamic Acid- Based Inhibitor 10O, PDB code: 4zx4:
Jump to Fluorine binding site number: 1; 2; 3;
In total 3 binding sites of Fluorine where determined in the X-Ray Crystal Structure of Pfa-M1 in Complex with Hydroxamic Acid- Based Inhibitor 10O, PDB code: 4zx4:
Jump to Fluorine binding site number: 1; 2; 3;
Fluorine binding site 1 out of 3 in 4zx4
Go back to
Fluorine binding site 1 out
of 3 in the X-Ray Crystal Structure of Pfa-M1 in Complex with Hydroxamic Acid- Based Inhibitor 10O
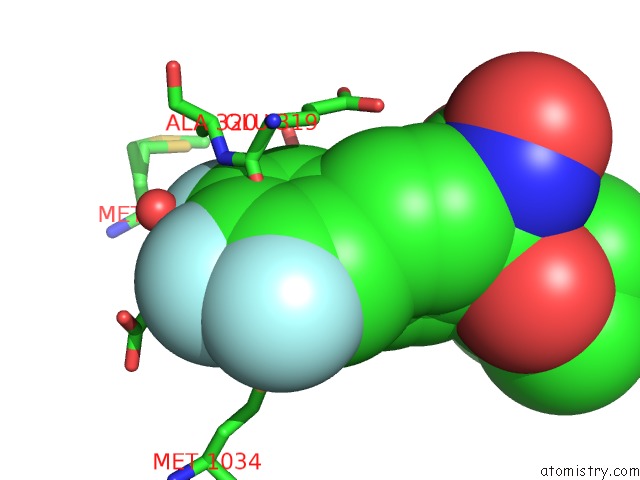
Mono view

Stereo pair view

Mono view

Stereo pair view
A full contact list of Fluorine with other atoms in the F binding
site number 1 of X-Ray Crystal Structure of Pfa-M1 in Complex with Hydroxamic Acid- Based Inhibitor 10O within 5.0Å range:
|
Fluorine binding site 2 out of 3 in 4zx4
Go back to
Fluorine binding site 2 out
of 3 in the X-Ray Crystal Structure of Pfa-M1 in Complex with Hydroxamic Acid- Based Inhibitor 10O

Mono view

Stereo pair view

Mono view

Stereo pair view
A full contact list of Fluorine with other atoms in the F binding
site number 2 of X-Ray Crystal Structure of Pfa-M1 in Complex with Hydroxamic Acid- Based Inhibitor 10O within 5.0Å range:
|
Fluorine binding site 3 out of 3 in 4zx4
Go back to
Fluorine binding site 3 out
of 3 in the X-Ray Crystal Structure of Pfa-M1 in Complex with Hydroxamic Acid- Based Inhibitor 10O

Mono view

Stereo pair view

Mono view

Stereo pair view
A full contact list of Fluorine with other atoms in the F binding
site number 3 of X-Ray Crystal Structure of Pfa-M1 in Complex with Hydroxamic Acid- Based Inhibitor 10O within 5.0Å range:
|
Reference:
N.Drinkwater,
N.B.Vinh,
S.N.Mistry,
R.S.Bamert,
C.Ruggeri,
J.P.Holleran,
S.Loganathan,
A.Paiardini,
S.A.Charman,
A.K.Powell,
V.M.Avery,
S.Mcgowan,
P.J.Scammells.
Potent Dual Inhibitors of Plasmodium Falciparum M1 and M17 Aminopeptidases Through Optimization of S1 Pocket Interactions. Eur.J.Med.Chem. V. 110 43 2016.
ISSN: ISSN 0223-5234
PubMed: 26807544
DOI: 10.1016/J.EJMECH.2016.01.015
Page generated: Thu Aug 1 07:27:55 2024
ISSN: ISSN 0223-5234
PubMed: 26807544
DOI: 10.1016/J.EJMECH.2016.01.015
Last articles
Cl in 5WGSCl in 5WGT
Cl in 5WGR
Cl in 5WG9
Cl in 5WFZ
Cl in 5WFQ
Cl in 5WFW
Cl in 5WFP
Cl in 5WFM
Cl in 5WEO